
Back ترسيب مناعي للكروماتين Arabic Hromatinska imunoprecipitacija BS Chromatin-Immunpräzipitation German Inmunoprecipitación de cromatina Spanish Immunoprécipitation de chromatine French מיצוי נוגדני של כרומטין HE Kromatin-immunprecipitáció Hungarian Immunoprecipitazione della cromatina Italian クロマチン免疫沈降 Japanese Chromatine-immunoprecipitatie Dutch
Chromatin immunoprecipitation (ChIP) is a type of immunoprecipitation experimental technique used to investigate the interaction between proteins and DNA in the cell. It aims to determine whether specific proteins are associated with specific genomic regions, such as transcription factors on promoters or other DNA binding sites, and possibly define cistromes. ChIP also aims to determine the specific location in the genome that various histone modifications are associated with, indicating the target of the histone modifiers.[1] ChIP is crucial for the advancements in the field of epigenomics and learning more about epigenetic phenomena.[2]
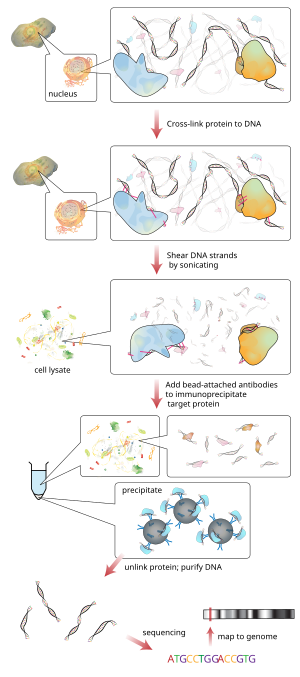
Briefly, the conventional method is as follows:
- DNA and associated proteins on chromatin in living cells or tissues are crosslinked (this step is omitted in Native ChIP).
- The DNA-protein complexes (chromatin-protein) are then sheared into ~500 bp DNA fragments by sonication or nuclease digestion.
- Cross-linked DNA fragments associated with the protein(s) of interest are selectively immunoprecipitated from the cell debris using an appropriate protein-specific antibody.
- The associated DNA fragments are purified and their sequence is determined. Enrichment of specific DNA sequences represents regions on the genome that the protein of interest is associated with in vivo.
- ^ Collas, Philippe. (January 2010). "The Current State of Chromatin Immunoprecipitation". Molecular Biotechnology. 45 (1): 87–100. doi:10.1007/s12033-009-9239-8. PMID 20077036. S2CID 24225210.
- ^ Rosenfeld, John M; Cooke, Tracy; Li, Zirong; Saito, Kan; Taganov, Konstantin; Thyagarajan, Bhaskar; Solache, Alejandra (March 2013). "Systematic optimization of parameters involved in preparation of chromatin and chromatin immunoprecipitation (ChIP) workflow". Epigenetics & Chromatin. 6 (S1): P122, 1756–8935–6-S1-P122. doi:10.1186/1756-8935-6-S1-P122. ISSN 1756-8935. PMC 3620580.